Molecular Biological Tools - MBTs
Molecular Biological Tools (MBTs) are analyses used to estimate biodegradation at contaminated sites. They can provide key evidence about contaminant-degrading microorganisms and biodegradation processes at many phases associated with site remediation projects. Here, we describe MBT fundamentals and introduce how the available MBTs work. Although numerous MBTs exist, project managers should be informed about a large swath of the approaches available so that the correct MBT can be selected that best align with site-specific project objectives and concerns that need to be addressed.
Related Article(s):
- Bioremediation - Anaerobic
- Metagenomics
- Proteomics and Proteogenomics
- Quantitative Polymerase Chain Reaction (qPCR)
- Stable Isotope Probing (SIP)
Contributor(s): Dora Ogles-Taggart and Dr. Brett Baldwin
Key Resource(s):
- Groundwater Sampling & Analysis Using qPCR[1]
- Guide for Assessing Biodegradation and Source Indentification of Groundwater Conmaninants Using CSIA[2]
Introduction
Molecular biological tools (MBTs) is a collective term for a group of laboratory analyses that are now commonly used to evaluate biodegradation potential or activity at contaminated sites. “Molecular” refers to the fact that the analyses are performed directly on cellular biomolecules including DNA, RNA, phospholipids, and proteins. “Biological’ refers to the application of the molecular tools to study biological activity including bioremediation processes. “Tools” include a variety of scientific methods and techniques that are commercially available. MBTs are also included in the more general term “environmental molecular diagnostics”.
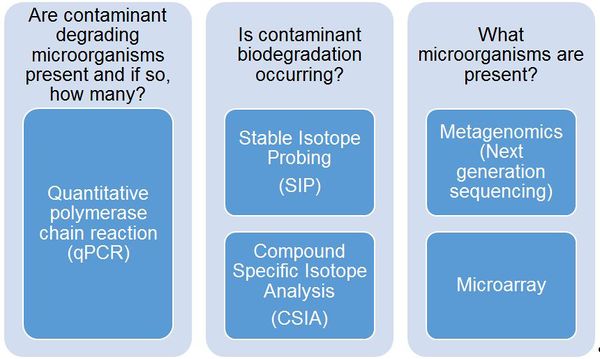
MBTs provide a crucial line of evidence during site characterization, remedy selection, and performance monitoring for answering specific questions related to contaminant biodegradation (Fig. 1).
Traditionally, techniques such as heterotrophic and “contaminant-specific” plate counts are used to assess the potential for in situ biodegradation at contaminated sites. However, often <1% of bacteria can be cultivated (grown) in the laboratory[3]. Thus, cultivation-based techniques can vastly underestimate targeted microbial populations and are generally not suitable to enumerate organisms of interest. An MBT, like quantitative polymerase chain reaction (qPCR), eliminates the biases of cultivation-based methods and thus provides more accurate assessment of microbial processes.
Available MBTs
Quantitative Polymerase Chain Reaction (qPCR)
Quantitative polymerase chain reaction (qPCR) is a DNA-based technique used to detect and quantify specific microorganisms or functional genes that can biodegrade contaminants of concern. qPCR is commonly used to support decisions regarding remedy selection, remedy design, and performance monitoring. For example, qPCR quantification of Dehalococcoides mccartyi and vinyl chloride reductase genes is indispensable at sites impacted by perchloroethylene (PCE), trichloroethylene (TCE) and other chlorinated solvents[1]. qPCR analysis is also commonly used to quantify functional genes involved in aerobic and anaerobic biodegradation of benzene, toluene, ethylbenzene, and xylenes (BTEX) to evaluate monitored natural attenuation (MNA) and other treatment strategies at petroleum hydrocarbon impacted sites[4][5][6][7]. See an article about qPCR here: Quantitative Polymerase Chain Reaction (qPCR)
Reverse transcriptase quantitative polymerase chain reaction (RT-qPCR) is a variation of qPCR [4]. RT-qPCR is based on the analysis of RNA, rather than DNA, to quantify gene expression and indicate biodegradation activity. In order to produce the enzymes that are biodegrading the contaminant, the genes must be expressed – transcribed from DNA into the corresponding mRNA sequence. Since RT-qPCR is based on analysis of RNA, the results reflect whether biodegradation is actively occurring.
QuantArrays are a hybrid technology that use very small reaction volumes, but in other respects are the same as conventional qPCR and RT-qPCR. The primary advantage of QuantArrays is the simultaneous and accurate quantification of a broad spectrum of target genes in a single analysis for more comprehensive evaluation of contaminant biodegradation. Simultaneous quantification of a suite of target genes is particularly useful at sites where there are mixtures of contaminants or multiple potential biodegradation pathways or treatment approaches are being evaluated.
Stable Isotope Probing (SIP)
Stable isotope probing (SIP) tracks the environmental fate of a 13C labeled contaminant to determine whether biodegradation of the contaminant of concern is occurring in situ. Essentially, the 13C label serves as a tracer. If biodegradation of the contaminant is occurring, the 13C label will be detected in the end products of biodegradation. More specifically, for compounds that serve as a carbon or energy source, the 13C label will be detected in the biomolecules (phospholipids, DNA, proteins) of contaminant degraders or mineralized to CO2.
In practice, SIP is most commonly used to determine if biodegradation of petroleum hydrocarbons (BTEX, PAHs) or oxygenates (MTBE, TBA) is occurring under existing site conditions to evaluate the feasibility of MNA as a site management strategy[8][9][10][11]. While less commonly employed in practice, SIP can also be used in conjunction with DNA based analyses to help identify the organisms involved in specific biodegradation processes[12][13][14].
Compound Specific Isotope Analysis (CSIA)
Compound specific isotope analysis (CSIA) is an analytical method that measures the ratio of stable isotopes (e.g., 13C/12C, 2H/1H) of a contaminant of concern (e.g., TCE). For some compounds, isotopic ratios change in predictable ways (e.g., isotopic fractionation) as the compound is degraded whereas physical processes like volatilization and dilution do not appreciably shift the ratio(s)[2]. For compounds where degradation leads to isotopic enrichment, CSIA provides information on whether biodegradation is occurring and on the extent of degradation. See an article about CSIA here: Compound Specific Isotope Analysis (CSIA)
Metagenomics (Next Generation DNA Sequencing)
Metagenomics is defined as the direct genetic analysis of collective genomes present in an environmental sample[15]. It includes techniques like clone libraries and more recently next generation DNA sequencing methods. Researchers most frequently use metagenomics in environmental remediation applications to assess biodiversity and investigate microbial community composition. For example, metagenomics analysis revealed that bacteria of the order Oceanospirillales and alkane degraders were enriched in the dissolved plume stemming from the Deepwater Horizon oil spill[16]. Outside of research settings, metagenomics analysis may be most applicable as an exploratory tool for investigating biodegradation of emerging contaminants where degrading microorganisms have not been isolated and biodegradation pathways have not yet been elucidated. See an article about metagenomics here: Metagenomics
Microarrays
Microarrays are comprised of thousands to more than a million short segments of DNA called probes that are used to detect the presence of corresponding target genes in an environmental sample. The most widely recognized microarrays in the industry are the PhyloChip and GeoChip[17]. As with metagenomics, PhyloChip analysis is most often used to examine biodiversity and shifts in microbial community composition in response to site activities. At a TCE impacted site at Ft. Lewis for example, PhyloChip analysis revealed a diverse microbial community in groundwater samples (over 1,300 operational taxonomic units [OTUs]) and increases in the relative abundances of Bacteroidetes, Firmicutes, δ-Proteobacteria, Chloroflexi suggesting stimulation of halorespiring and fermenting bacteria following electron donor addition[18]. PhyloChip and GeoChip have also been used to investigate changes in microbial community composition and function at radionuclide sites[19][20][21] and following the Deepwater Horizon oil spill[22].
Summary
MBTs provide a crucial line of evidence during site characterization, remedy selection, and performance monitoring. Although each MBT provides valuable information, site managers should select a MBT based on the site-specific questions that need to be addressed (e.g., Fig. 1).
References
- ^ 1.0 1.1 Lebrón, C. A., Dennis, P., Acheson, C., Barros, N., Major, D., Petrovskis, E., Loffler, F. E., Ritalahti, K. M., Yeager, C. M., Edwards, E. A., Hatt, J. K. and Ogles, D. M., 2014. Standardized procedures for use of nucleic acid-based tools - Recommendations for groundwater sampling and analysis using qPCR. ER-1561. Strategic Environmental Research Development Program, Arlington, VA. ER-1561
- ^ 2.0 2.1 Hunkeler, D., Meckenstock, R. U., Sherwood Lollar, B., Schmidt, T. C. and Wilson, J. T., 2008. A Guide for Assessing Biodegradation and Source Identification of Organic Groundwater Contaminants Using Compound Specific Isotope Analysis (CSIA). U.S. Environmental Protection Agency, Washington, D.C., EPA/600/R-08/148. Report pdf
- ^ Amann, R.I., Ludwig, W., Schleifer, K.H., 1995. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiological Reviews, 59(1), 143-169. Article
- ^ 4.0 4.1 Baldwin, B.R., Biernacki, A., Blair, J., Purchase, M.P., Baker, J.M., Sublette, K., Davis, G., Ogles, D., 2010. Monitoring gene expression to evaluate oxygen infusion at a gasoline-contaminated site. Environmental Science & Technology, 44(17), 6829-6834. doi:10.1021/es101356t
- ^ Baldwin, B.R., Nakatsu, C.H. and Nies, L., 2003. Detection and enumeration of aromatic oxygenase genes by multiplex and real-time PCR. Applied and Environmental Microbiology, 69(6), 3350-3358. doi:10.1128/AEM.69.6.3350-3358.2003
- ^ Beller, H.R., Kane, S.R., Legler, T.C., Alvarez, P.J., 2002. A real-time polymerase chain reaction method for monitoring anaerobic, hydrocarbon-degrading bacteria based on a catabolic gene. Environmental Science & Technology, 36(18), 3977-3984. doi:10.1021/es025556w
- ^ DeBruyn, J.M., Chewning, C.S., Sayler, G.S., 2007. Comparative quantitative prevalence of Mycobacteria and functionally abundant nidA, nahAc, and nagAc dioxygenase genes in coal tar contaminated sediments. Environmental science & technology, 41(15), 5426-5432. doi:10.1021/es070406c
- ^ Busch‐Harris, J., Sublette, K., Roberts, K.P., Landrum, C., Peacock, A.D., Davis, G., Ogles, D., Holmes, W.E., Harris, D., Ota, C.,Yang, X., 2008. Bio‐Traps Coupled with Molecular Biological Methods and Stable Isotope Probing Demonstrate the In Situ Biodegradation Potential of MTBE and TBA in Gasoline‐Contaminated Aquifers. Groundwater Monitoring & Remediation, 28(4), 47-62. doi:10.1111/j.1745-6592.2008.00216.x
- ^ Geyer, R., Peacock, A.D., Miltner, A., Richnow, H.H., White, D.C., Sublette, K.L., Kästner, M., 2005. In situ assessment of biodegradation potential using biotraps amended with 13C-labeled benzene or toluene. Environmental Science & Technology, 39(13), 4983-4989. doi:10.1021/es048037x
- ^ Key, K.C., Sublette, K.L., Johannes, T.W., Ogles, D., Baldwin, B., Biernacki, A., 2014. Assessing BTEX Biodegradation Potential at a Refinery Using Molecular Biological Tools. Groundwater Monitoring & Remediation, 34(1), 35-48. doi:10.1111/gwmr.12037
- ^ Williams, N., Hyland, A., Mitchener, R., Sublette, K., Key, K.C., Davis, G., Ogles, D., Baldwin, B., Biernacki, A., 2013. Demonstrating the In Situ Biodegradation Potential of Phenol Using Bio‐Sep® Bio‐Traps® and Stable Isotope Probing. Remediation Journal, 23(1), 7-22. doi:10.1002/rem.21335
- ^ Aslett, D., Haas, J. and Hyman, M., 2011. Identification of tertiary butyl alcohol (TBA)-utilizing organisms in BioGAC reactors using 13C-DNA stable isotope probing. Biodegradation, 22(5), 961-972. doi:10.1007/s10532-011-9455-3
- ^ Hatzinger, P., Fuller, M., 2014. New approaches to evaluate the biological degradation of RDX in groundwater. Project ER-1607. Strategic Environmental Research Development Program, Arlington, VA. ER-1607
- ^ Key, K.C., Sublette, K.L., Duncan, K., Mackay, D.M., Scow, K.M., Ogles, D., 2013. Using DNA‐Stable Isotope Probing to Identify MTBE‐and TBA‐Degrading Microorganisms in Contaminated Groundwater. Groundwater Monitoring & Remediation, 33(4), 57-68. doi:10.1111/gwmr.12031
- ^ Stenuit, B., Eyers, L., Schuler, L., Agathos, S.N., George, I., 2008. Emerging high-throughput approaches to analyze bioremediation of sites contaminated with hazardous and/or recalcitrant wastes. Biotechnology Advances, 26(6), 561-575. doi: 10.1016/j.biotechadv.2008.07.004
- ^ Mason, O.U., Hazen, T.C., Borglin, S., Chain, P.S., Dubinsky, E.A., Fortney, J.L., Han, J., Holman, H.Y.N., Hultman, J., Lamendella, R., Mackelprang, R., 2012. Metagenome, metatranscriptome and single-cell sequencing reveal microbial response to Deepwater Horizon oil spill. The ISME Journal, 6(9), 1715-1727. doi:10.1038/ismej.2012.59
- ^ Hazen, T.C., Rocha, A.M., Techtmann, S.M., 2013. Advances in monitoring environmental microbes. Current Opinion in Biotechnology, 24(3), 526-533. doi:10.1016/j.copbio.2012.10.020
- ^ Lee, P.K., Warnecke, F., Brodie, E.L., Macbeth, T.W., Conrad, M.E., Andersen, G.L., Alvarez-Cohen, L., 2011. Phylogenetic microarray analysis of a microbial community performing reductive dechlorination at a TCE-contaminated site. Environmental Science & Technology, 46(2), 1044-1054. doi:10.1021/es203005k
- ^ Brodie, E.L., DeSantis, T.Z., Joyner, D.C., Baek, S.M., Larsen, J.T., Andersen, G.L., Hazen, T.C., Richardson, P.M., Herman, D.J., Tokunaga, T.K., Wan, J.M., 2006. Application of a high-density oligonucleotide microarray approach to study bacterial population dynamics during uranium reduction and reoxidation. Applied and Environmental Microbiology, 72(9), 6288-6298. doi:10.1128/AEM.00246-06
- ^ Chakraborty, R., Wu, C.H. and Hazen, T.C., 2012. Systems biology approach to bioremediation. Current Opinion in Biotechnology, 23(3), 483-490. doi:10.1016/j.copbio.2012.01.015
- ^ Xu, M., Wu, W.M., Wu, L., He, Z., Van Nostrand, J.D., Deng, Y., Luo, J., Carley, J., Ginder-Vogel, M., Gentry, T.J., Gu, B., 2010. Responses of microbial community functional structures to pilot-scale uranium in situ bioremediation. The ISME Journal, 4(8), 1060-1070. doi:10.1038/ismej.2010.31
- ^ Hazen, T.C., Dubinsky, E.A., DeSantis, T.Z., Andersen, G.L., Piceno, Y.M., Singh, N., Jansson, J.K., Probst, A., Borglin, S.E., Fortney, J.L., Stringfellow, W.T., Bill, M., Conrad, M. E., Tom, L. M., Chavarria, K. L., Alusi, T. R., Lamendella, R., Joyner, D. C., Spier, C., Baelum, J., Auer, M., Zemla, M. L., Chakraborty, R., Sonnenthal, E. L., D’haeseleer, P., Holman, H.-Y. N., Osman, S., Lu, Z., Van Nostrand, J. D., Deng, Y., Zhou, J., Mason, O. U., 2010. Deep-sea oil plume enriches indigenous oil-degrading bacteria. Science, 330(6001), 204-208. doi: 10.1126/science.1195979
See Also
- Environmental Molecular Diagnostics
- Compound Specific Isotope Analysis: The Science, Technology and Selected Examples from the Literature with Application to Fuel Oxygenates and Chlorinated Solvents
- SERDP and ESTCP Expert Panel Workshop on Research and Development Needs for the Environmental Remediation Application of Molecular Biological Tools
- Cryogenic Collection of Complete Subsurface Samples for Molecular Biological Analysis
- Impacts of Sampling and Handling Procedures on DNA- and RNA-Based Microbial Characterization and Quantification
- Standardized Procedures for Use of Nucleic Acid-Based Tools
- Prokaryotic cDNA Subtraction: A Method to Rapidly Identify Functional Gene Biomarkers
- BioReD: Biomarkers and Tools for Reductive Dechlorination Site Assessment, Monitoring, and Management
- Application of Microarrays and qPCR to Identify Phylogenetic and Functional Biomarkers Diagnostic of Microbial Communities that Biodegrade Chlorinated Solvents to Ethene
- Molecular Biomarkers for Detecting, Monitoring, and Quantifying Reductive Microbial Processes
- Quantifying the Presence and Activity of Aerobic, Vinyl Chloride-Degrading Microorganisms in Dilute Groundwater Plumes by Using Real-Time PCR
- Application of Nucleic Acid-Based Tools for Monitoring MNA, Biostimulation and Bioaugmentation at Chlorinated Solvent Sites
- Online Lecture Course - Molecular Evidence for Biodegradation